What does bisulfite sequencing detect?
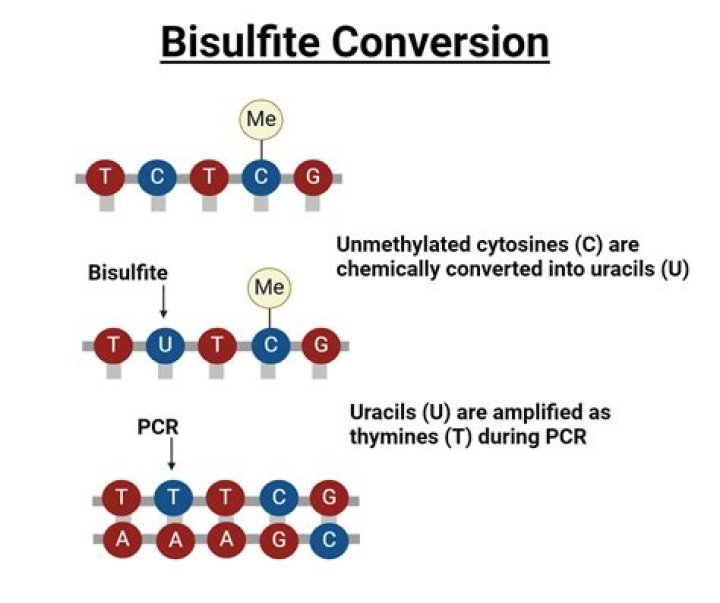
Bisulfite sequencing (also known as bisulphite sequencing) is the use of bisulfite treatment of DNA before routine sequencing to determine the pattern of methylation. DNA methylation was the first discovered epigenetic mark, and remains the most studied.
What information can you get from a ChIP-seq experiment?
ChIP-seq is primarily used to determine how transcription factors and other chromatin-associated proteins influence phenotype-affecting mechanisms. Determining how proteins interact with DNA to regulate gene expression is essential for fully understanding many biological processes and disease states.
What is BS seq?
Bisulfite sequencing (BS-Seq) or whole-genome bisulfite sequencing (WGBS) is a well-established protocol to detect methylated cytosines in genomic DNA . In this method, genomic DNA is treated with sodium bisulfite and then sequenced, providing single-base resolution of methylated cytosines in the genome.
What is the difference between ChIP and ChIP-seq?
For instance, ChIP-seq generally produces profiles with higher spatial resolution, dynamic range, and genomic coverage, allowing it to have higher sensitivity and specificity over ChIP-chip in terms of protein binding site identification.
How expensive is bisulfite sequencing?
Pricelist
| Bulk Epigenomics Library Preparation | Internal (WCM, WCM Qatar & Cornell U) |
|---|---|
| WGBS (Whole Genome Bisulfite Sequencing) | $250 |
| ChIP-seq | $150 |
| ATACseq (library prep from cells) | $225 |
| ATACseq partial (price per sample) | $100 |
What is the formula of bisulfite?
Sodium bisulfite
| PubChem CID | 23665763 |
|---|---|
| Chemical Safety | Laboratory Chemical Safety Summary (LCSS) Datasheet |
| Molecular Formula | NaHSO3 or NaHO3S or HNaO3S |
| Synonyms | SODIUM BISULFITE 7631-90-5 Sodium hydrogen sulfite sodium hydrogensulfite Sodium bisulphite More… |
| Molecular Weight | 104.06 |
What is ChIP analysis?
Chromatin immunoprecipitation, or ChIP, is an antibody-based technology used to selectively enrich specific DNA-binding proteins along with their DNA targets. ChIP is used to investigate a particular protein-DNA interaction, several protein-DNA interactions, or interactions across the whole genome or a subset of genes.
What is a ChIP to ChIP experiment?
ChIP-on-chip (also known as ChIP-chip) is a technology that combines chromatin immunoprecipitation (‘ChIP’) with DNA microarray (“chip”). The goal of ChIP-on-chip is to locate protein binding sites that may help identify functional elements in the genome.
What is hi c sequencing?
Hi-C sequencing is high‐throughput chromosome conformation capture technique to analyze spatial genome organization and map higher‐order chromosome folding and topological associated domains. Hi-C allows quantification of interactions between all possible pairs of DNA fragments simultaneously.
How much does a DNA methylation test cost?
This is the most popular method for methylation profiling, which sits between whole genome bisulfite sequencing and low throughput methods that can access the methylation of a single locus. Over 360 publications to date used Illumina methylation arrays. According to Illumina, the price is about U.S. $300–360/sample.
What is high-throughput bisulfite sequencing?
Bisulfite sequencing is the gold-standard for measuring methylation over the genomes of interest. Here, we review several techniques used for the analysis of high-throughput bisulfite sequencing.
What is reduced representation bisulfite sequencing (RRBS)?
Reduced representation bisulfite sequencing (RRBS) is another technique that can also profile DNA methylation at single-base resolution. It combines digestion of genomic DNA with restriction enzymes and sequencing with bisulfite treatment in order to enrich for areas with high CpG content.
What is whole genome bisulfite sequencing (WGBS)?
Whole genome bisulfite sequencing (WGBS) is considered the ‘gold standard’ for assaying DNA methylation because it provides global coverage at single-base resolution. Briefly, it combines bisulfite conversion of DNA molecules with high-throughput sequencing.