What is gene expression normalization?
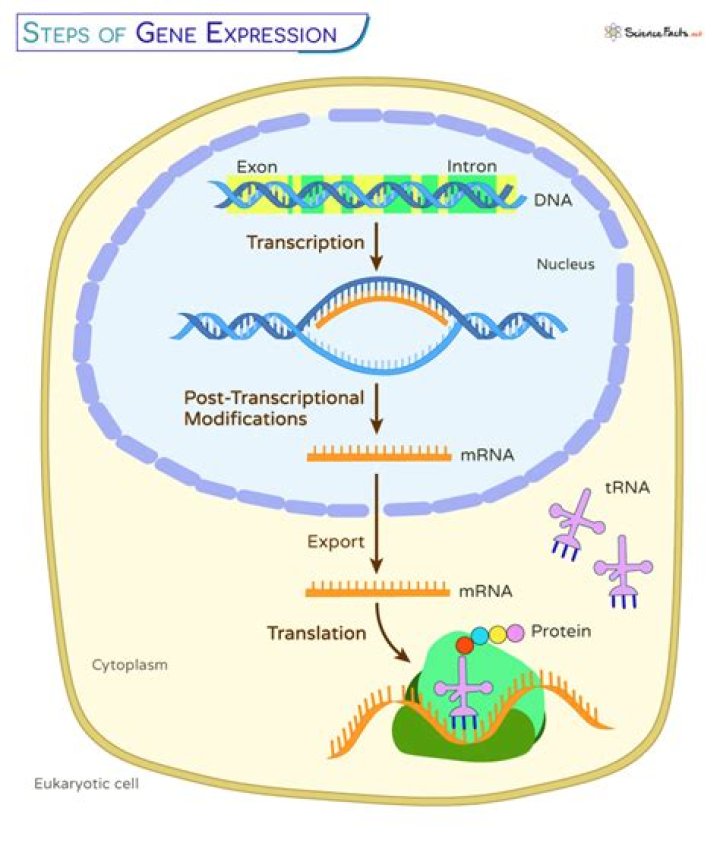
Normalization is achieved by dividing expression values by the total intensity (i.e., the sum of all expression values) of the given array. Centralization11 assumes that regulation is well behaved, i.e., most genes are not significantly regulated or about equal numbers of genes are up- and down-regulated.
What is FPKM expression?
FPKM stands for Fragments Per Kilobase of transcript per Million mapped reads. In RNA-Seq, the relative expression of a transcript is proportional to the number of cDNA fragments that originate from it.
How does TMM normalization work?
TMM normalization is a simple and effective method for estimating relative RNA production levels from RNA-seq data. The TMM method estimates scale factors between samples that can be incorporated into currently used statistical methods for DE analysis.
Why is gene expression data normalized?
Raw gene expression data from these high-throughput technologies must be normalized to remove technical variation so that meaningful biological comparisons can be made. Data normalization is a crucial step in the gene expression analysis as it ensures the validity of its downstream analyses (Lovén et al., 2012).
How do you choose normalization method?
The best normalization technique is one that empirically works well, so try new ideas if you think they’ll work well on your feature distribution….Summary.
| Normalization Technique | Formula | When to Use |
|---|---|---|
| Clipping | if x > max, then x’ = max. if x < min, then x’ = min | When the feature contains some extreme outliers. |
How do you convert FPKM to RPKM?
With paired-end reads RPKM = 2 x FPKM.
What is meant by normalization of data in DBMS?
Normalization is the process of organizing data in a database. This includes creating tables and establishing relationships between those tables according to rules designed both to protect the data and to make the database more flexible by eliminating redundancy and inconsistent dependency.
How do you normalize TPM?
Count up all the RPK values in a sample and divide this number by 1,000,000. This is your “per million” scaling factor. Divide the RPK values by the “per million” scaling factor. This gives you TPM.”
Is there an easy-to-use platform for isoform expression analysis?
While analysis of isoform expression is important for discovering tumor-specific isoform signatures and interpreting relevant genomic mutations, there is currently no web-based, easy-to-use, and publicly available platform for this purpose.
What is RNA-Seq normalization?
RNA-Seq normalization explained. Published on November 28, 2016. RNA-Seq (short for RNA sequencing) is a type of experiment that lets us measure gene expression. The sequencing step produces a large number (tens of millions) of cDNA1 fragment sequences called reads. Every read represents a part of some RNA molecule in the sample2.
Is the Red isoform more expressed in the first experiment?
If we rely on raw counts, the first experiment has produced more red fragments, and so we may conclude that the red isoform is more expressed there. Computing R P K won’t change anything since we are comparing expression of the same isoform.
Where can I find the isoexpresso database?
The ISOexpresso database is available online at Alternative splicing occurs in most human genes with multiple exons as a regulatory process of gene expression [ 1 ].