What is the rate-limiting enzyme of purine synthesis?
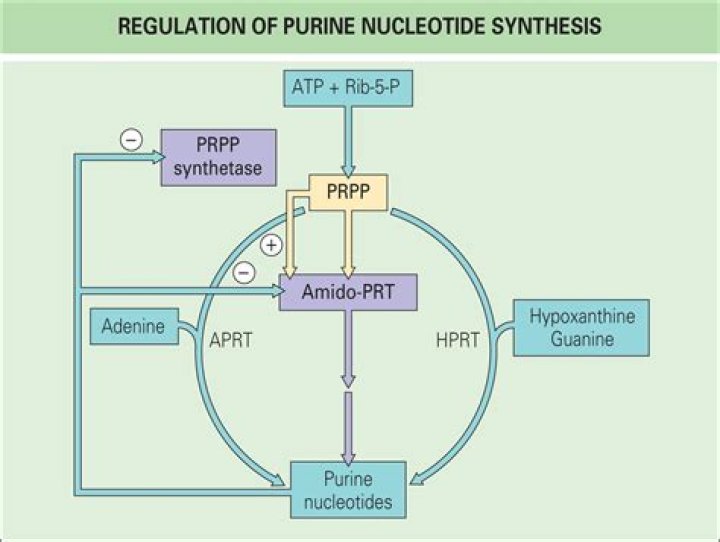
Amido phosph-oribosyltransferase (ATase) catalyzes the first step of the de novo purine synthesis pathway, the conversion of PRPP into 5-phosphoribosylamine. ATase is the rate-limiting enzyme in the de novo purine synthesis pathway.
What is the limiting substance for the synthesis of purines?
PRPP
In Step 1, ribose-5-phosphate is activated via the direct transfer of a pyrophosphoryl group from ATP to C-1 of the ribose, yielding 5-phosphoribosyl-a-pyrophosphate (PRPP) (Figure 27.3). The enzyme is ribose-5-phosphate pyrophosphokinase. PRPP is the limiting substance in purine biosynthesis.
What is the rate-limiting step in pyrimidine synthesis?
The conversion of IMP to XMP is the rate-limiting step in the de novo synthesis of guanine nucleotides and is catalyzed by IMP dehydrogenase (IMPDH or PUR13; EC 1.1. 1.205).
How is AMP synthesized from IMP?
IMP is the major branch-point in the formation of all purine nucleotides. In one branch, IMP is irreversibly converted into AMP in two enzymatic steps. First, succinyl-AMP (sAMP) is formed by condensation of aspartic acid and IMP catalyzed by adenylosuccinate synthase. This reaction is driven by GTP hydrolysis.
What inhibits purine synthesis?
Also Methotrexate indirectly inhibits purine synthesis by blocking the metabolism of folic acid (it is an inhibitor of the dihydrofolate reductase). Allopurinol is a drug that inhibits the enzyme xanthine oxidoreductase and, thus, lowers the level of uric acid in the body.
Which is the rate limiting step of pyrimidine synthesis that exhibits allosteric inhibition by cytidine triphosphate?
Aspartate transcarbamoylase step
Aspartate transcarbamoylase step—The second main source of pyrimidine ring aspartate combines with carbamoyl phosphate in the presence of aspartate transcarbamoylase. This step is the committed step of the pathway as this enzyme is allosterically regulated (allosteric inhibition by CTP). 3.
What is the rate limiting step in pyrimidine synthesis that exhibits allosteric inhibition by cytidine triphosphate?
Aspartate transcarbamoylase step—The second main source of pyrimidine ring aspartate combines with carbamoyl phosphate in the presence of aspartate transcarbamoylase. This step is the committed step of the pathway as this enzyme is allosterically regulated (allosteric inhibition by CTP). 3.
Where are purines synthesized?
the liver
Purine synthesis occurs in all tissues. The major site of purine synthesis is in the liver and, to a limited extent, in the brain.
How is purine synthesis regulated?
Purine nucleotide biosynthesis is regulated at several steps IMP synthesis is controlled by the levels of adenine and guanine nucleotides. Additional control is exerted by feedforward activation, which is the stimulation of a subsequent enzyme by the preceding substrate.
How many enzymes are involved in de novo purine biosynthesis?
In humans, this metabolic transformation is carried out in ten steps by sequential orchestration of six enzymes. The first reaction in the de novo purine biosynthetic pathway is the conversion of PRPP to 5-phosphoribosylamine (PRA) by PRPP amidotransferase (PPAT) and is presumed to be rate-limiting.
Is the purinosome a novel level of metabolic organization of enzymes?
This review highlights the purinosome as a novel level of metabolic organization of enzymes in cells, its consequences for regulation of purine metabolism, and the extent that purine metabolism is being targeted for the treatment of cancers. Keywords: purine metabolism, metabolon, purinosome
How much ATP is required for purine synthesis?
The de novo purine synthesis pathway requires several moles of ATP for generation of each mole of purine nucleotide product, while HPRT and APRT require one ATP.
What is the activator and initiator molecule for purine synthesis?
The activator molecule for Purine synthesis is PRPP, which activates the enzyme AmidoPhospho Ribosyl transferase The initiator molecule for the synthesis is Ribose-5-Phosphate. The optimum concentration of Ribos-5-Phosphate is maintained by the enzyme Glucose-6-Phosphate dehydrogenase, which is the regulatory enzyme of Hexose Mono Phosphate Shunt.