What is the substrate of serine proteases?
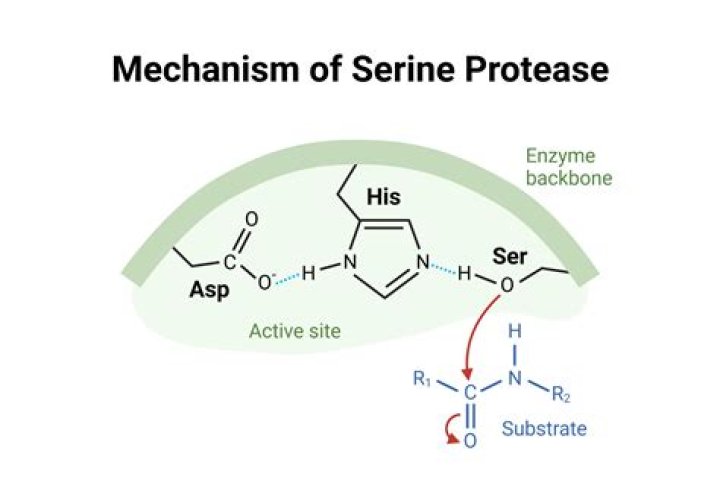
Serine proteases are enzymes that cleave peptide bonds in proteins, in which serine serves as the nucleophilic amino acid at the active site….Introduction.
| Proteinase | Sources | Substrates/activities |
|---|---|---|
| Cathepsin G | P Monocytes | |
| Proteinase-3 | Plasma proteins (immunoglobulins, clotting factors, complement components) |
How are serine proteases activated?
It is activated by cleavage through trypsin. As can be seen, trypsinogen activation to trypsin is essential, because it activates its own reaction, as well as the reaction of both chymotrypsin and elastase. Therefore, it is essential that this activation does not occur prematurely.
What determines substrate specificity of proteolytic enzymes?
Substrate specificity of a protease is determined by molecular interactions at the protein-protein interface of protease and substrate in the binding cleft of the protease. Amino acid side chains of the substrate are accommodated within subpockets of the protease.
Do all serine proteases have a catalytic triad?
All the mammalian serine proteases have a common 3D structure, although there are quite significant differences in some regions of the surface, reflecting the different physiological functions, and hence need for different interactions with different molecules. They all have the same catalytic mechanism.
Where does serine protease cleave?
Serine proteases are usually endoproteases and catalyze bond hydrolysis in the middle of a polypeptide chain. However, several families of exoproteases have been described that remove one or more amino acids from the termini of target polypeptide chains.
What determines the specificity of chymotrypsin?
The Active Site Environment A specific pocket adjacent to the active site triad determines the specificity of the protease (chymotrypsin cleaves adjacent to large aromatic side chains, trypsin adjacent to lys or arg residues). These are amino acids 189, 216 and 226 which line a pocket adjacent to the active site triad.
Where do serine proteases cleave?
How is substrate specificity determined?
The traditional method used to investigate the reaction specificity of an enzyme with different substrates is to perform individual kinetic measurements. The multiplex ESI-MS assay is an accurate and valid method for substrate specificity evaluation, in which multiple substrates can be evaluated in one assay.
What does serine do in the catalytic triad?
The serine esterases have a catalytic triad: serine, glutamic or aspartic acid, and histidine. These catalytic residues are responsible for the nucleophilic attack of the active site serine on the carbonyl carbon atom of the ester.
Which of the following is serine proteases?
Clan PA proteases bearing the trypsin fold are the largest family of serine proteases and perhaps the best studied group of enzymes. Digestive enzymes such as trypsin and chymotrypsin cleave polypeptide chains at positively charged (Arg/Lys) or large hydrophobic (Phe/Trp/Tyr) residues, respectively.
How many proteolytic enzymes are serine?
Over one third of all known proteolytic enzymes are serine proteases. Among these, the trypsins underwent the most predominant genetic expansion yielding the enzymes responsible for digestion, blood coagulation, fibrinolysis, development, fertilization, apoptosis, and immunity.
What is the catalytic triad of serine proteases?
Serine proteases perform their catalytic roles using three key residues, which are commonly referred to as the catalytic triad: Ser, His, Asp. Highlight the chymotrypsin catalytic triad. The elements are color coded as follows: C, O, N .
What is the difference between trypsin and elastase binding pockets?
The trypsin binding pocket contains Asp189 to select for positively charged sidechains, such as arginine. The arginine sidechain is part of a larger peptide-based inhibitor called aeruginosin 98-B, which is now shown in balls and sticks. The elastase binding pocket is more constrained, explaining the preference for smaller residues.