Where does the enzyme HindIII cut?
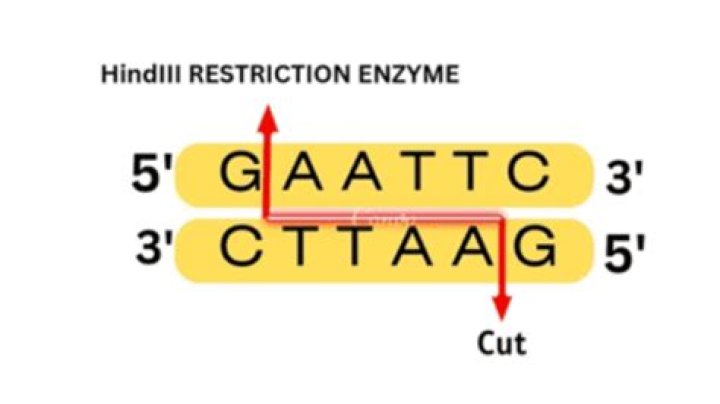
Thermo Scientific HindIII restriction enzyme recognizes A^AGCTT sites and cuts best at 37°C in R buffer. See Reaction Conditions for Restriction Enzymes for a table of enzyme activity, conditions for double digestion, and heat inactivation for this and other restriction enzymes.
What sequence is cut by Hind 2?
Hind II recognizes the sequence GTPy/PuAC and generates fragments with blunt ends (1). Compatible ends Hind II generates fragments with blunt ends and is compatible to any other blunt end. Isoschizomers Hind II is an isoschizomer to Hinc II. Hind II is inhibited by 6-methyladenine as indicated (*).
Where do Type 2 restriction enzymes cut?
Type II restriction endonucleases always cleave at or near their recognition sites. They produce small, well-defined fragments of DNA that help to characterize genes and genomes and that produce recombinant DNAs. Fragments of DNA produced by restriction endonucleases can be moved from one organism to…
What are the two types of restriction enzyme cuts?
Today, scientists recognize three categories of restriction enzymes: type I, which recognize specific DNA sequences but make their cut at seemingly random sites that can be as far as 1,000 base pairs away from the recognition site; type II, which recognize and cut directly within the recognition site; and type III.
Does Hind 3 produce blunt ends?
Option B: Hind 3: It is a type 2 restriction endonuclease which gives sticky ends. It is isolated from Haemophilus influenzae. Eco RV: It is type 2 endonuclease producing blunt ends in the centre of nucleotide sequence GAT/ATC.
What is the source of Hind 2?
HindIII (pronounced “Hin D Three”) is a type II site-specific deoxyribonuclease restriction enzyme isolated from Haemophilus influenzae that cleaves the DNA palindromic sequence AAGCTT in the presence of the cofactor Mg2+ via hydrolysis.
Is HindIII sticky or blunt?
Recognition Sequences
| Enzyme | Organism | Blunt or Sticky End |
|---|---|---|
| HindIII | Haemophilus influenzae Rd | Sticky |
| Hinfl | Haemophilus influenzae Rf | Sticky |
| Sau3A | Staphylococcus aureus | Sticky |
| AluI | Arthrobacter luteus | Blunt |
What is the application of hind III enzyme in research?
Uses in research. HindIII as well as other type II restriction endonucleases are very useful in modern science, particularly in DNA sequencing and mapping. Unlike type I restriction enzymes, type II restriction endonucleases perform very specific cleaving of DNA.
What is the function of NDE I?
In molecular biology, it is commonly used as a restriction enzyme . The ends generated by Nde I digest: Nde I is a specific Type II restriction enzyme that cuts open specific target sequences, unlike exonucleases.
What is the best temperature for ligation of NDE I-generated ends?
Ligation of Nde I-generated ends is therefore best performed with higher ligase concentration with a longer ligation time, whether at room temperature, 14-16 ° C, or at 4 ° C. ^ Watson RJ, Schildraut I, Qiang BQ, Martin SM, Visentin LP (Dec 13, 1982).
What is the function of Ndende I?
Nde I is a specific Type II restriction enzyme that cuts open specific target sequences, unlike exonucleases. This enzyme is used in gene cloning to cut open reading frames in the plasmid of certain bacteria such as E. coli and insert a foreign gene, such as the gfpuv gene that codes for bio fluorescence of the jelly fish Aequorea victoria .